Congratulations to Dr. Kamal El-Din Hani for receiving the State Incentive Award in the field of agricultural sciences .
Announcement to activate the electronic course Infectious Diseases 2
Congratulations to Prof. Dr. Abdul Muhaimin Mustafa for his appointment as Head of the Department of Anatomy and Embryology
Announcement for students of the second level, the fourth level, the examination of the work of the year, a subject of guidance and veterinary information

Announcement of the year's work exam, first level anatomy of birds and fish

Exam schedule of the year's work

Distribution schedule for the year's work exams for all academic levels
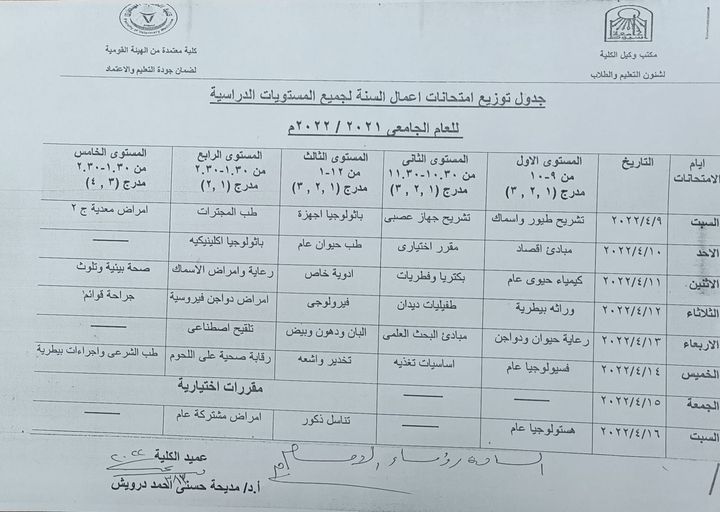
elective courses
Molecular Variation between RT-PCR Detected Rotavirus Infection of Naturally Diarrheic Neonatal Calves and Rotavirus Strains of Commercial Vaccines
Neonatal diarrhea is the main cause of morbidity and mortality in calves, and Rotavirus is the main viral etiology.
Rotavirus vaccines are one of the main important methods for control of diarrhea in neonates’ calves. In the current
study, Deoxyribonucleic acid (DNA) sequencing and phylogenetic analysis of Bovine Rotavirus Group a (BRVA)
were performed in our study. 1 Calf guard® vaccine genotype (G6P1) and 5 different field genotypes (2 G6P5, 1
G10P5, G10P? and 1 G10P11) were subjected to DNA sequencing. We observed that at the nucleotide level, G10P5
and G10P? Sequences were 100 % identical with each other, two G6P5 sequences were 100% identical with each
other and there was no significant similarity between sequences of G10P11 with sequences of G6P5, G10P5, and
G10P? The phylogenetic analysis of G10P5 and G10P? Isolates showed a close cluster with G10 isolates of Sharkia
governorate, Egypt, phylogenetic analysis of two G6P5 and one G10P11 isolate showed a close cluster with the VP4
gene of Rotavirus isolates of Dakahlia Governorate, Egypt. Molecular comparison between detected and typed Rotaviruses’
genotypes with other genotypes of common vaccines indicated that there were genetically close or distance
between field and vaccine Rotavirus strains. Our results can be concluded as the following, Molecular comparison
between detected and typed Rotaviruses’ genotypes with other genotypes of common vaccines indicated that there
was genetically close or distance between field and vaccinal Rotavirus strains. Also, we suggest that Rotavac vaccine
containing G6P5 Rotavirus strain and Scour guard vaccine containing can be used in Assiut governorate due to circulating
of G6P5 and G10P11 strains of Rotavirus in Assiut.
Comprehensive quantitation of multi-signature peptides originating from casein for the discrimination of milk from eight different animal species using LC-HRMS with stable isotope labeled peptides
Milk species adulteration has become an altering issue worldwide. In this study, a robust quantification method based on LC-HRMS for the simultaneous detection and differentiation of milk type from eight different animal species (namely: cow, water buffalo, wild yak, goat, sheep, donkey, horse, and camel) was established by detecting nine signature peptides originating from casein. The developed method was in-house validated in terms of sensitivity, accuracy, and precision. As a result, limits of quantification (LOQ) were ranging from 5 to 30 µg/L, recoveries ranged from 95.2% to 104.5%, and intra-day and inter-day variability were lower than 11.4% and 12.6%, respectively, for all the targeted peptides. Furthermore, this method was successfully applied to 46 commercial minor species’ milk, in which 15 samples were false labeling. The obtained results indicate the necessity to monitor milk species adulteration in order to protect consumers from consuming misleading labeled minor species animal’s milk.



